I spent some time recently working on a small side project that I’m excited to share with you all today. It’s an A/B test significance calculator and you can check it out at ABTestCalculator.com.
What’s an A/B test significance calculator, you ask? At a high level, A/B testing is a technique that allows you to improve your website by showing visitors one of several versions of something and then measuring the impact each version has on some other event. For example, you might A/B test the wording of a button on your homepage and find that it increases the number of people who sign up by 10%. An A/B test significance calculator helps you analyze the results of an A/B test to determine whether there is a statistically significant change that is not just the result of random chance.
The math is somewhat complicated which is why a number of A/B test calculators exist, including isvalid.org by Evan Solomon, another by KISSmetrics, another by VWO, and many more.
Why build another? Three reasons: to learn the math, to get better at JavaScript, and to build a tool that makes the results of an A/B test easier to understand.
I think most of these other tools do users a disservice by not clearly explaining how to interpret the results. They tend to throw around the percentage improvement and significance figures without explaining what they mean which in the past has led me to make uninformed and often wrong decisions. Worse, most don’t tell you when you don’t have enough participants or conversions in your test and will happily apply statistical analysis to your results even when those methods don’t apply.
It is my hope with this tool that users leave with a clearer understand of how to interpret the results. A few features:
- An executive summary that provides an overview in plain English about how to interpret the results
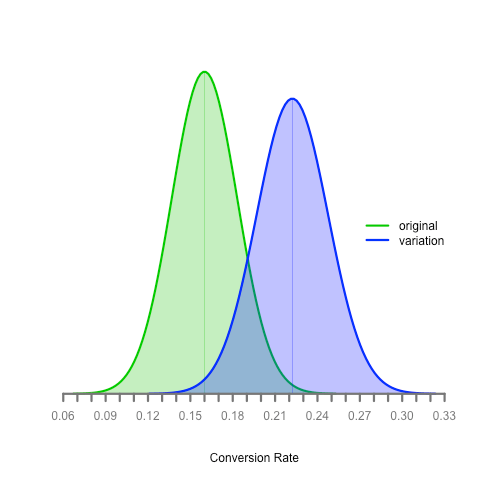
- One graph showing where the true conversion rate for each variation falls (using something called a Wald approximation) and another showing the percentage change between those two distributions
- It handles ties as well as tests where there aren’t enough participants or conversions to come to a conclusion
- Results are significant when there is at least a 90% chance that one variation will lead to an improvement
- The ability to copy a URL for the results to make them easier to share
If you have any suggestions on how to make it better please don’t hesitate to let me know.
On the coding site of things, most of the JavaScript I’ve done in the past (including Preceden and Lean Domain Search) has been with lots and lots of messy jQuery. A lot of new JavaScript technologies have come out in the last few years and I was put on a project at Automattic not too long ago that uses many of them. I fumbled around with it, getting stuff done but not really understanding the fundamentals.
I’m happy to say that this tool uses React for the view layer, NPM and Browserify for dependency management, Gulp for its build system, parts of ES6 for the code (courtesy of Babel), JSHint for code analysis, Mocha for testing, and Github Pages for hosting — all of which I had little to no experience with when I started this project. If you’re interested in checking it out, all of the code is open source (my first!) so you can view it on Github.
This project is the best JavaScript I know how to do so if you do check out the code, please let me know if you have any suggestions on how to improve it.
One last note in case you were wondering about the domain: the former owner had a simple A/B test calculator up on it, but wasn’t actively working on it so I found his email via WHOIS, offered him $200 for it, he countered with $700, I countered with $300 and we had a deal. Normally I wouldn’t pay someone for a domain (I heard there is this amazing service to help people find available domains…), but the price was right and I figured it was worth it for the credibility and SEO value it adds. When I showed him the new site recently all he responded with was “I’m pretty glad I sold the domain now!” which was nice :).
Thanks for checking it out!