In yesterday’s post, we showed that a binomial distribution can be approximated by a normal distribution and some of the math behind it.
Today we’ll take it a step further, showing how those results can help us understand the distribution of a sample proportion.
Consider the following example:
Out of the last 250 visitors to your website, 40 signed up for an account.
The conversion rate for that group is 16%, but it’s possible (and likely!) that the true conversion rate differs from this. Statistics can help us determine a range (a confidence interval) for what the true conversion rate actually is.
Recall that in the last post we said that the mean of the binomial distribution can be approximated with a normal distribution with a mean and standard deviation calculated by:
For a proportion, we want to figure out the mean and standard deviation on a per-trial basis so we divide each formula by n, the number of trials:
With the mean and standard deviation of the sample proportion in hand, we can plot the distribution for this example:
As you can see, the most likely conversion rate is 16% (which is no surprise), but the true conversion rate can fall anywhere under that curve with it being less and less likely as you move farther away.
Where it gets really interesting is when you want to compare multiple proportions.
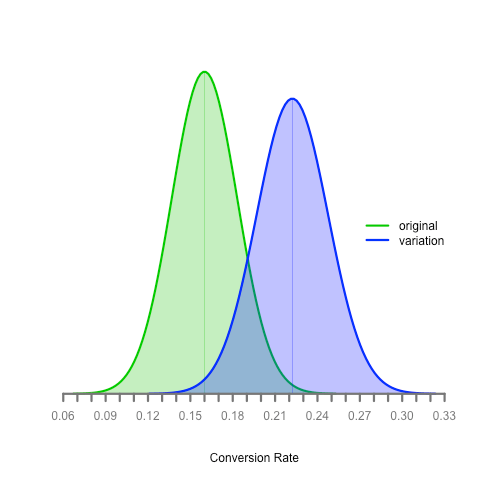
Let’s say we’re running an A/B test and the original had 40 conversions out of 250 like the example above and the experimental version had 60 conversions out of 270 participants. We can plot the distribution of both sample proportions with this code:
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| setClass( | |
| Class = "Distribution", | |
| representation = representation( | |
| name = "character", | |
| participants = "numeric", | |
| conversions = "numeric", | |
| sample_proportion = "numeric", | |
| se = "numeric", | |
| color = "character", | |
| x = "vector", | |
| y = "vector" | |
| ) | |
| ) | |
| # We rewrite the initialize method for Distribution objects so that we can | |
| # set the x and y values which are used throughout the plotting process | |
| setMethod( | |
| f = "initialize", | |
| signature = "Distribution", | |
| definition = function( .Object, name, participants, conversions, color ) { | |
| .Object@name = name | |
| .Object@sample_proportion = conversions / participants | |
| .Object@se = sqrt( ( .Object@sample_proportion * ( 1 – .Object@sample_proportion ) ) / participants ) | |
| .Object@color = color | |
| .Object@x = seq( -4, 4, length = 100 ) * .Object@se + .Object@sample_proportion | |
| .Object@y = dnorm( .Object@x, .Object@sample_proportion, .Object@se ) | |
| return ( .Object ) | |
| } | |
| ) | |
| # Given a list of distributions, this returns a list of the x and y axis range | |
| get_axis_ranges = function( distributions ) { | |
| x_all = vector() | |
| y_all = vector() | |
| for ( distribution in distributions ) { | |
| x_all = c( x_all, distribution@x ) | |
| y_all = c( y_all, distribution@y ) | |
| } | |
| xlim = c( min( x_all ), max( x_all ) ) | |
| ylim = c( min( y_all ), max( y_all ) ) | |
| # Note that by forming a list of the vectors, the vectors get converted to lists | |
| # which we then have to convert back to vectors in order to use them for plotting | |
| return ( list( xlim, ylim ) ) | |
| } | |
| get_x_axis_values = function( x_range ) { | |
| by = 0.01 | |
| min_x = floor( min( x_range ) / by ) * by | |
| max_x = ceiling( max( x_range ) / by ) * by | |
| return ( seq( min_x, max_x, by = by ) ) | |
| } | |
| # Define the distributions that we want to plot | |
| distributions = list( | |
| new( Class = "Distribution", name = "original", participants = 250, conversions = 40, color = "#00cc00" ), | |
| new( Class = "Distribution", name = "variation", participants = 270, conversions = 60, color = "blue" ) | |
| ) | |
| # Determine the range to use for each axis | |
| axis_range = get_axis_ranges( distributions ) | |
| xlim = axis_range[[ 1 ]] | |
| ylim = axis_range[[ 2 ]] | |
| # Create the plot | |
| plot( NULL, NULL, type = "n", xlim = xlim, ylim = ylim, xlab = "Conversion Rate", ylab = "", main = "", axes = FALSE ) | |
| # Render each of the curves | |
| line_width = 3 | |
| for ( distribution in distributions ) { | |
| polygon( distribution@x, distribution@y, col = adjustcolor( distribution@color, alpha.f = 0.3 ), border = NA ) | |
| lines( distribution@x, distribution@y, col = adjustcolor( distribution@color, alpha.f = 1 ), lwd = line_width ) | |
| # Draw a line down the center of the curve | |
| ci_center_y = dnorm( distribution@sample_proportion, distribution@sample_proportion, distribution@se ) | |
| coords = xy.coords( c( distribution@sample_proportion, distribution@sample_proportion ), c( 0, ci_center_y ) ) | |
| lines( coords, col = adjustcolor( distribution@color, alpha.f = 0.4 ), lwd = 1 ) | |
| } | |
| # Render the x axis | |
| axis( side = 1, at = get_x_axis_values( xlim ), pos = 0, col = "#777777", col.axis = "#777777", lwd = line_width ) | |
| # Finally, render a legend | |
| legend_text = vector() | |
| legend_colors = vector() | |
| for ( distribution in distributions ) { | |
| legend_text = c( legend_text, distribution@name ) | |
| legend_colors = c( legend_colors, distribution@color ) | |
| } | |
| legend('right', legend_text, lty = 1, lwd = line_width, col = legend_colors, bty = 'n' ) |
Here’s the result:
What can we determine from this? Is the experimental version better than the original? What if the true proportion for the original is 20% (towards the upper end of its distribution) and the true proportion for the experimental version is 16% (towards the lower end of its distribution)?
We’ll save the answer for a future post :)